 |
Ph.D., Distinguished Professor Department of Biomedical Informatics
Stony Brook Cancer Center
Stony Brook, NY 11794
Phone: (631) 638-2590
Email: Ramana.Davuluri@stonybrookmedicine.edu |
|
INTERESTS
Machine Learning; Molecular Data Science; Foundation Models for Genomics; DNABERT; Bioinformatics; Cancer Genomics
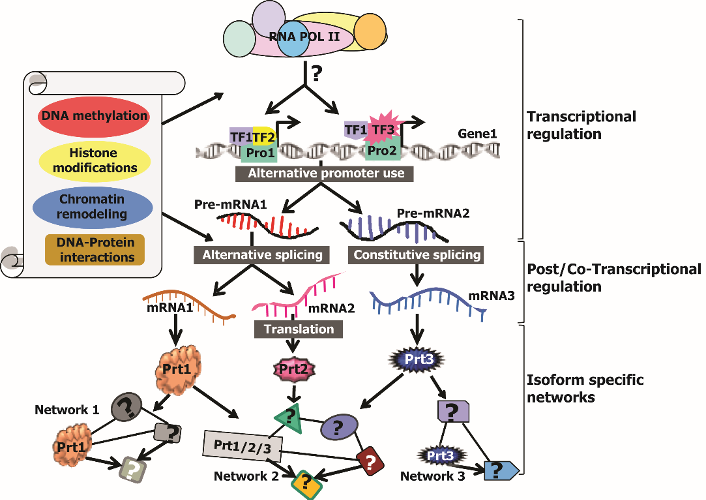
Summary of Ongoing Research Projects:
Developing novel deep-learning based methods for deciphering non-coding gene regulatory code (1R01LM013722): To develop novel pre-trained DNA Bidirectional Encoder Representations from Transformers, called DNABERT, and associated deep-learning tools to decipher the language of non-coding DNA and facilitate integration of gene regulatory information from rapidly accumulating sequence data with NLM’s genetic databases (for example, dbSNP, dbGaP and ClinVar), which serve both scientists and the public health by helping identify the genetic components of disease. The goal of our proposed research is to develop DNABERT for a variety of sequence prediction tasks, and benchmark with existing state-of-the-art deep-learning based methods. Specific aims are (1) develop novel deep-learning methods by adapting BERT; (2) apply the proposed deep-learning methods to specifically target non-coding DNA sequence analyses and predictions; and (3) predict and validate functional non-coding genetic variants by applying DNABERT prediction models. A major contribution of the proposed research is development of pre-trained DNABERT model and prediction algorithms, which present new powerful methods for analyses and predictions of DNA sequences.
Molecular signaling in mechanobiology regulation by single-cell analyses using bioinformatics approach (NSF – PI: Yi-Xian Qin, co-PI: Davuluri) : The project will develop a single-cell multiplex in situ tagging (scMIST) system combined with advanced machine leaning algorithms through successive rounds of labeling and imaging to effectively achieve a multiplexity of thousands of data points using a common fluorescence microscope and a simple procedure in a typical biological laboratory setting, which has the potential to revolutionize the field of mechanical biology. Davuluri group provides Bioinformatics and Machine-Learning expertise to the research project.
The spinal cell atlas of opioid-targeted inflammasomes in the HIV pain model: mechanism and pathogenic roles (NIDA/NIH; 1R01DA062257; PI: Tang, Shao Jun; Co-I: Davuluri) : Davuluri, as co-Investigator, provides bioinformatics support for analysis of NGS data that will be generated by this project.
DNABERT for interpretation of germline and somatic non-coding mutations in cancer : This project aims to develop (1) novel deep-learning methods, by adapting DNABERT, to identify combinations of regulatory SNPs that are associated with cancer risk and survival, and (2) software for region selection and prediction models that demonstrate the utility of genome-wide DNABERT-mv based mutational profiles for cancer prediction tasks. Variant data from large AllOfUs cohort will be used for training the models, while TCGA, ICGC and other cancer data portals will be used for testing the models.
DNABERT: pre-trained Bidirectional Encoder Representations from Transformers model for DNA-language in genome: Understanding the hidden instructions within genome on gene regulation is crucial for biological research. However, complex language patterns widely exist in DNA, such as polysemy and distant semantic relationship, which previous methods often fail to capture especially in data-scarce scenarios. For the first time, Davuluri group (in collaboration with Dr Han Liu, Department of Computer Science, Northwestern University) is developing DNABERT to form global understanding of genomic sequences based on up and downstream sequence contexts. Using an innovative global contextual embedding of input sequences, DNABERT attempts to tackle the problem of sequence specificity prediction with a “top-down” approach by developing general understanding of DNA language via self-supervised pre-training and applying it to specific tasks (for example, prediction of promoters, transcription factor binding sites and splice sites), in contrast to the traditional “bottom-up” approach using task-specific data. Various modules of DNABERT are currently under development. It is anticipated that the pre-trained DNABERT on the human genome can also be readily applied to data from other organisms with exceptional performance.
Algorithms and bioinformatics software for promoter prediction: As a postdoctoral fellow in Michael Zhang group, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, Davuluri developed novel algorithms and computer programs for predicting gene-promoters and non-coding first exons, a highly complex problem that remained as a critical gap in gene-prediction for several years. These creative and groundbreaking approaches facilitated the prediction and annotation of Pol-II promoters in human and mouse genomes.
Data-mining methods for integrative analysis of transcriptome and epigenome data: In collaboration with Dr. Tim Huang group at Ohio State University Comprehensive Cancer Center, Davuluri performed integrative microarray technology and statistical modeling approaches to predict which proteins work with estrogen to contribute to breast cancer development. The computational predictions in this study indicated that the interaction of estrogen with one of seven different partner proteins determines whether the gene is activated or suppressed in breast cancer cells. This was a noteworthy data-mining methodology breakthrough because it allowed integrative analysis of big datasets, often consisting of expression, chromatin landscaping data and TF-binding information for thousands of genomic loci, for predicting small networks of transcription factors, which then could be validated by more traditional experimental biology techniques. This is one of the early integrative chromatin immunoprecipitation (ChIP)-chip/ChIP-seq and gene expression profiling studies, which laid foundations for efficient methods for integrative analysis of multi-omics data on a genomic scale.
Isoform-level gene expression and regulation in mammalian development and cancer: Recent genome-wide studies have discovered that majority of human genes produce multiple transcript-variants/protein-isoforms, which could be involved in different functional pathways. Moreover, altered expression of specific isoforms for numerous genes is linked with cancer and its prognosis, as cancer cells manipulate regulatory mechanisms to express specific isoforms that confer drug resistance and survival advantages. For example, cancer-associated alterations in alternative exons and splicing machinery have been identified in cancer samples, suggesting that specific transcript-variants could be more effective as diagnostic and prognostic markers than corresponding genes. In a recent study, Davuluri group discovered that majority of genes associated with neurological diseases expressed multiple transcripts through alternative promoters by using integrative NextGen sequencing based experimental approaches and bioinformatics analysis. The study also observed aberrant use of alternative promoters and splice variants in different cancers. Subsequently, his group demonstrated that cancer cell-lines regardless of their tissue of origin can be effectively discriminated from non-cancer cell-lines at isoform-level, but not at gene-level. The novel informatics methods have been successfully applied by his collaborators in different cancer studies.
Platform-independent isoform-level gene signatures for stratification of cancer patients into molecular subgroups: Based on recent studies from Davuluri group and others, significant expression differences were observed between different sample groups (e.g., developmental stages, cancer subtypes, normal vs cancer) for numerous genes at the isoform-level but not at the overall gene-level. Davuluri group investigated whether the isoform-level transcriptome changes could provide better patient stratification in terms of overall prognosis and classification accuracy. His group developed novel methods, by integrating data discretization, feature selection, and meta-classification algorithms, for derivation of platform-independent gene signature for multi-label molecular stratification of cancer patients, from exon-array and RNA-seq data. The application of these algorithms has led to the development of new methods for diagnosis of glioblastoma and other cancers and investigation of alternative splicing on drug-target gene interactions.
Algorithms and bioinformatics software for analyses of NextGen sequence data: Mapping genome-wide data to human subtelomeres has been problematic due to the incomplete assembly and challenges of lowcopy repetitive DNA elements. Davuluri group developed novel bioinformatics pipelines for incorporating multi-read mapping for annotation of the updated assemblies using short-read data sets from ChIP-seq data, and RNA-seq data. As part of other collaborative efforts, his group developed bioinformatics methods for identification of single-nucleotide polymorphisms (SNPs) that alter miRNA gene regulation and influence tumor susceptibility. Similarly, his group played a pivotal role in the development of informatics methods required for analysis of small-RNA sequence data, with Nishikura group at Wistar Institute, Philadelphia, PA.
Selected publications:
- Surana P, Dutta P, Davuluri RV (2024). TransTEx: Novel tissue-specificity scoring method for grouping human transcriptome into different expression groups. Bioinformatics, 40 (8).
- Zhou Z, Wu W, Ho H, Wang J, Shi L, RV Davuluri RV, Wang Z, Liu H (2024) Dnabert-s: Learning species-aware dna embedding with genome foundation models, ArXiv.
- Zhou Z, Ji Y, Li W, Dutta P, Davuluri RV, Liu H. (2024) Dnabert-2: Efficient foundation model and benchmark for multi-species genome. Proceedings of ICLR 2024; May 7–11, 2024; Vienna Austria2024 (in the proceedings of the twelfth International Conference on Learning Representations)..
- Ji Y, Dutta P, Davuluri RV. (2023). Deep Multi-Omics Integration by Learning Correlation-Maximizing Representation Identifies Prognostically Better Stratified Cancer Subtypes. Bioinformatics Advances, 3(1):vbad075.
- Keathley R, Kocherginsky M, Davuluri R, Matei D. (2023). Integrated Multi-Omic Analysis Reveals Immunosuppressive Phenotype Associated with Poor Outcomes in High-Grade Serous Ovarian Cancer. Caners (Basel), 15(14):3649.
- Yang L, Dutta P, Davuluri R, Wang J. (2023) Rapid, High-Throughput Single-Cell Multiplex In Situ Tagging (MIST) Analysis of Immunological Disease with Machine Learning. Anal Chem. 95(19):7779-7787.
- Y Ji, X Tong, DD Xu, J Liao, Davuluri RV, Yang GY, Mishra RK. (2023). A new robust classifier to detect hot-spots and null-spots in protein–protein interface: validation of binding pocket and identification of inhibitors in in vitro and in vivo models. Big Data Analytics in Chemoinformatics and Bioinformatics, 247-263.
- Ji Y, Zhou Z, Liu H, Davuluri RV. (2021) DNABERT: pre-trained Bidirectional Encoder Representations from Transformers model for DNA-language in genome. Bioinformatics. Epub 2021/02/05. doi: 10.1093/bioinformatics/btab083. PubMed PMID: 33538820.
- Chakraborty A, Ay F, Davuluri RV. (2021) ExTraMapper: Exon- and Transcript-level mappings for orthologous gene pairs. Bioinformatics. Epub 2021/05/21. doi: 10.1093/bioinformatics/btab393. PubMed PMID: 34014317.
- Ji Y, Mishra R, Davuluri RV (2020) In silico analysis of alternative splicing on drug target genes. Sci Rep. 10(1):134.
- Shilpi A, Kandpal M, Ji Y, Seagle BL, Shahabi S, Davuluri RV (2019) Platform-independent classification system for predicting high-grade serous ovarian carcinoma molecular subtypes. JCO Clin Cancer Inform, 3, 1-9.
- Dapas M, Kandpal M, Bi Y, Davuluri RV (2017). Comparative evaluation of isoform-level gene expression estimation algorithms for RNA-seq and exon-array platforms. Briefings in Bioinformatics Mar 1;18(2):260-269. doi: 10.1093/bib/bbw016.
- Jin H-J, Jung S, DebRoy A, Davuluri RV (2016). Identification and validation of regulatory SNPs that modulate transcription factor chromatin binding and gene expression in prostate cancer Oncotarget 7:54616-54626, doi: 10.18632/oncotarget.10520.
- Jung S, Bi Y, Davuluri RV (2015). Evaluation of data discretization methods to derive platform independent isoform expression signatures for multi-class tumor subtyping. BMC Genomics. Nov 10;16 Suppl 11:S3. doi: 10.1186/1471-2164-16-S11-S3. PMCID: PMC4652565.
- Pal S, Bi Y, Macyszyn L, Showe LC, O’Rourke DM and Davuluri RV. (2014) Isoform-level gene signature improves prognostic stratification and accurately classifies glioblastoma subtypes. Nucleic Acids Res. Epub 2014/02/08. doi: 10.1093/nar/gku121. PubMed PMID: 24503249.
- Bi Y and Davuluri RV (2013) NPEBseq: Nonparametric Empirical Bayesian based Procedure for Differential Expression Analysis from RNA-seq Data. BMC Bioinformatics, 14: 262. Highly accessed.
- Ota, H.*, Sakurai, M.*, Gupta, R.*, Valente, L., Wulff, B.-E., Ariyoshi, K., Iizasa, H., Davuluri, R.V. and Nishikura, K. (2013) ADAR1 complexes with Dicer and plays a role in microRNA processing and RNA-induced gene silencing mechanisms. Cell, 153(3): 575-589. (*Equal contribution)
- Zhang, Z., Pal, S., Tchou, J. and Davuluri, R.V. (2013) Isoform-level expression profiles provide better cancer signatures than gene-level expression profiles, Genome Medicine, Apr 17;5(4):33.
- Pal, S., Gupta, R., Kim, H., Wickramasinghe, P., Baubet, V., Showe, L.C., Dahmane, N. and Davuluri, R.V. (2011) Alternative transcription exceeds alternative splicing in generating the transcriptome diversity of cerebellar development, Genome Research, 21(8): 1260-1272.
- Davuluri R.V., Grosse, I. and Zhang, M.Q. (2001). Computational identification of first exons and promoters in the human genome. Nature Genetics, 29: 412-417. (Note: This work was featured in Nature Reviews Genetics, 3: 3-9; in Bioinformatics section of Highlights as “Filling the gap in gene prediction” January, 2002).
Present Lab Members & Trainees
Past Trainees in my Research Laboratory
Rotation Students and CSIRE Program Trainees
Davuluri has trained 20 graduate students and postdocs, and 4 junior investigators so far. These include that have gone on to faculty positions in academia (Sharmistha Pal, Scientist, Dana Farber Cancer Institute, Harvard University, Boston, MA; Hao Sun, Associate Professor, The Chinese University of Hong Kong, Hong Kong; Victor Jin, Professor, Dept of Molecular Medicine, UTHSCSA, San Antonio, TX) or leadership positions in industry or academia (Yingtao Bi, Bioinformatics Director, AbbVie Inc, Boston).
Course Director and Teaching:
- Course Director, BMI 511 Translational Bioinformatics